Scientists developed a new way to help understand what happens in the body when people consume a plant product and the many chemicals it contains. The American Chemical Society’s Journal of Natural Products published the method to quickly analyze the effects of a natural product, developed at Emory University.
Monday, March 16, 2026
Turning Over a New Leaf in Analyses of Natural Products
Scientists developed a new way to help understand what happens in the body when people consume a plant product and the many chemicals it contains. The American Chemical Society’s Journal of Natural Products published the method to quickly analyze the effects of a natural product, developed at Emory University.
Wednesday, January 7, 2026
Unlocking design secrets of deep-sea microbes
The microbe Pyrodictium abyssi is an archaeaon — a member of what’s known as the third domain of life — and an extremophile. It lives in deep-sea thermal vents, at temperatures above the boiling point of water, without light or oxygen, withstanding the enormous pressure at ocean depths of thousands of meters.
A biomatrix of tiny tubes of protein, known as cannulae, link cells of Pyrodictium abyssi together into a highly stable microbial community. No one knew how these single-celled microbes accomplished this feat of extreme engineering — until now.
A study using advanced microscopy techniques reveals new details about the elegant design of the cannulae and the remarkable simplicity of their method of construction. Nature Communications published the work, led by scientists at Emory University; the University of Virginia, Charlottesville; and Vrije Universiteit Brussel in Belgium.
The discovery holds the potential to inspire innovations in biotechnology, from the development of new “smart” materials to nanoscale drug delivery systems.
“Not only are the cannulae strong enough to endure extreme conditions, they’re beautiful,” says Vincent Conticello, Emory professor of chemistry and co-senior author of the paper. “To me, they resemble columns from the classical architecture of ancient Greece or Rome,” he adds, citing their fluted edges and precise regularity.
Related:
Tuesday, September 23, 2025
New methods expand access to molecules key to human health
Tuesday, April 15, 2025
New AI tool set to speed quest for advanced superconductors
The study was led by theorists at Emory University and experimentalists at Yale University. Senior authors include Fang Liu and Yao Wang, assistant professors in Emory’s Department of Chemistry, and Yu He, assistant professor in Yale’s Department of Applied Physics.
The team applied machine-learning techniques to detect clear spectral signals that indicate phase transitions in quantum materials — systems where electrons are strongly entangled. These materials are notoriously difficult to model with traditional physics because of their unpredictable fluctuations.
“Our method gives a fast and accurate snapshot of a very complex phase transition, at virtually no cost,” says Xu Chen, the study’s first author and an Emory PhD student in chemistry. “We hope this can dramatically speed up discoveries in the field of superconductivity.”
One of the challenges in applying machine learning to quantum materials is the lack of sufficient high-quality experimental data needed to train models. To overcome this, the researchers used high-throughput simulations to generate large amounts of data. They then combined these simulation results with just a small amount of experimental data to create a powerful and efficient machine-learning framework.
Read more about the discovery.
Related:
Monday, April 7, 2025
Chatbot opens computational chemistry to nonexperts
Advanced computational software is streamlining quantum chemistry research by automating many of the processes of running molecular simulations. The complicated design of these software packages, however, often limits their use to theoretical chemists trained in specialized computing techniques.
Wednesday, March 5, 2025
Atlanta Science Festival set to entertain, inspire and engage all ages
By Carol Clark
Atlanta Science Festival returns March 8-22, with more than 100 events throughout the metro area, inviting the public to join fun, interactive and educational experiences. The acclaimed city-wide celebration, one of the largest of its kind in the country, showcases the myriad science, technology engineering and mathematics (STEM) innovations happening in Atlanta, including at Emory.Tuesday, February 11, 2025
Plant extract inspires new chemistry and new early lead against triple-negative breast cancer
Chemists at Emory University invented a reaction to streamline the total synthesis of a compound, phaeocaulisin A, extracted from a plant used for centuries in traditional Chinese medicine.
Wednesday, November 13, 2024
Chemists showcase power of pathbreaking method to make complex molecules
Chemists synthesized a highly complex natural molecule through a revolutionary strategy of functionalizing normally inert carbon-hydrogen (C-H). Science published the breakthrough led by chemists at Emory University and Caltech.
Tuesday, October 29, 2024
Bacterial pathogen shows alarming resistance to common cleaners, chemists discover
Friday, March 29, 2024
A new estimate of U.S. soil organic carbon to improve Earth system models
By Carol Clark
Soil contains about twice as much carbon as the atmosphere and plants combined. It is a major carbon sink, capable of absorbing more carbon dioxide from the atmosphere than it releases. Management of soil carbon is key in efforts to mitigate climate change, in addition to being vital to soil health and agricultural productivity.Wednesday, March 6, 2024
Atlanta Science Festival returns to inspire discovery for all ages
Tuesday, December 12, 2023
New tool to analyze blood platelets holds major medical potential
By Carol Clark
A novel technique to test platelet function within a person’s blood sample is faster, easier and more precise than methods currently in use, an experimental study shows.Thursday, November 9, 2023
New antimicrobial shuts down bacterial growth without harming human cells
By Carol Clark
Scientists have shown how a molecule with broad-spectrum antibiotic activity works by disabling a process vital to bacterial growth without affecting the normal functioning of human cells. mBio, a journal of the American Society for Microbiology, published the work, led by researchers at Emory University and Pennsylvania State University.
The molecule, known as KKL-55, is one of a suite of recently identified molecules that interfere with a key bacterial mechanism known as trans-translation, essentially shutting down the ability of bacteria to grow.
“We’re opening a promising pathway for the development of new antibiotics to treat drug-resistant infections,” says Christine Dunham, co-corresponding author of the paper and a professor in Emory’s Department of Chemistry and the Emory Antibiotic Resistance Center.
Kenneth Keiler, a professor in the Department of Biochemistry and Molecular Biology at Pennsylvania State, is co-corresponding author of the paper.
First authors are Ha An Nguyen, who did the work as an Emory chemistry PhD candidate and has since graduated and works at Memorial Sloan Kettering, and Neeraja Marathe, a graduate student at Pennsylvania State.
A growing global threat
Antimicrobial-resistant infections have long been a public health threat. The situation grew even worse during the COVID-19 pandemic with increased antibiotic use and less prevention actions, according to the U.S. Centers for Disease Control and Prevention (CDC).
The CDC estimates that at least 2.8 million antimicrobial-resistant infections continue to occur in the United States each year, killing more than 35,000 people. Globally, the World Health Organization projects that these infections will cause up to 10 million deaths annually by 2050 if new antibiotics are not developed.
While antibiotics can save lives, any time they are used they can also contribute to the problem of resistance. Bacteria keep evolving new weapons as a defense against drugs, even as scientists work on developing new strategies to disarm bacteria.
Cross-toxicity, or harmful effects on humans, is another key drawback of some of the drugs used in a last-ditch effort to kill antibiotic-resistant bacteria.
Avoiding cross-toxicity
Dunham and Keiler are avoiding the problem of cross-toxicity by focusing on the inhibition of a mechanism unique to bacteria — trans-translation. This mechanism is vital to the proper functioning of the bacterial ribosome.
Keiler, a molecular geneticist and biochemist, first identified trans-translation in bacteria and is an expert in how it functions. Dunham, a structural biologist, is an expert in the human ribosome. She uses advanced biochemistry and structural biology techniques to understand the mechanics of molecular interactions.
“Our individual areas of expertise mesh well for this project,” Dunham says. “By collaborating, we are able to take the science further, faster.”
A cellular protein factory
The ribosome is an elaborate macromolecular machine within a cell that operates like a factory to manufacture proteins. Proteins are the machines that make cells run while nucleic acids such as DNA and RNA store the blueprints for life. The ribosome is made mostly of RNA, which does not just store information but can also act as an enzyme, catalyzing chemical reactions.
In a human cell, messenger RNA (mRNA), containing the instructions for manufacturing a protein, originates in the nucleus. While still in the nucleus, mRNA undergoes an elaborate quality-control process. It must pass inspection before getting exported to translate the information it contains into a protein.
“A lot of mRNAs have defects,” Dunham says. “Human cells have efficient ways to test mRNAs and ultimately remove the defective ones.”
Bacterial cells, however, have no nucleus or organized center for quality control.
“Bacteria wants to grow, grow and grow, which requires the ribosome to make a lot of proteins,” Dunham says. “But when mRNA has defects, there is little to no quality control. When the ribosome encounters a defective mRNA protein, synthesis gets stalled.”
The trans-translation process “rescues” ribosomes stalled due to such defects, in order to maintain proper protein synthesis and cell viability in bacteria.
How KKL-55 works
Using a high-throughput screening process, the Keiler lab has identified dozens of molecules that inhibit trans-translation in bacteria.
For the current paper, the researchers focused on understanding how one of these molecules, KKL-55, performs this trick. They used the high-powered structural biology technique of X-ray crystallography to capture KKL-55 in action as it interacted with a protein required for translation.
The results showed how KKL-55 blocks trans-translation by binding to elongation factor thermos-unstable (EF-Tu). EF-Tu is a protein that interacts with transfer RNA molecules, which play a key role in protein synthesis, and also transfer-messenger RNA, an RNA molecule required for the trans-translation pathway.
“We got lucky,” Dunham says. “There are dozens of steps involved in the process that KKL-55 could have inhibited and we might have had to test for each one. But the results are clear-cut. It shuts down trans-translation right at the beginning by preventing EF-Tu from binding to tmRNA.”
Determining the mechanism by which a molecule works to inhibit bacteria is a critical step toward developing a new antibiotic for clinical use. The next step is to test the efficacy of KKL-55 to treat a bacterial infection in a mouse model.
In 2021, the research team published their finding that a group of trans-translation inhibitors known as acylaminooxadiazoles clear multiple-drug-resistant Neisseria gonorrhoeae infection in mice after a single oral dose. That work is now advancing to clinical trials.
Dozens more trans-translation inhibitors await the team’s investigation. Each represents a potential new weapon to help humans stay on top in the arms race with drug-resistant bacteria.
Co-authors of the current paper include Alexandra Nagy (a former National Institutes of Health FIRST Institutional Research and Academic Career Development postdoctoral fellow at Emory who is now at Earlham College); as well as John Alumasa and Michael Vazquez (both from Pennsylvania State University). The work was funded by the National Institutes of Health, the National Institute of General Medical Sciences.
Related:
Images of enzymes in action reveal secrets of antibiotic-resistant bacteria
Wednesday, September 13, 2023
Natural compound found in plants inhibits deadly fungi
By Carol Clark
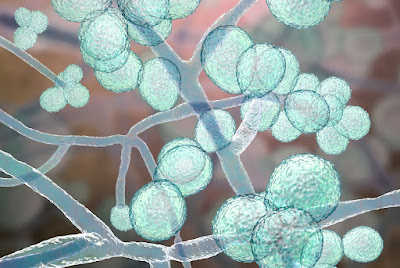
A new study finds that a natural compound found in many plants inhibits the growth of drug-resistant Candida fungi — including its most virulent species, Candida auris, an emerging global health threat. The journal ACS Infectious Diseases published the discovery led by scientists at Emory University.
Laboratory-dish experiments showed that the natural compound, a water-soluble tannin known as PGG, blocks 90% of the growth in four different species of Candida fungi. The researchers also discovered how PGG inhibits the growth: It grabs up iron molecules, essentially starving the fungi of an essential nutrient. By starving the fungi rather than attacking it, the PGG mechanism does not promote the development of further drug resistance, unlike existing antifungal medications.
Laboratory-dish experiments also showed minimal toxicity of PGG to human cells.
“Drug-resistant fungal infections are a growing healthcare problem but there are few new antifungals in the drug-development pipeline,” says Cassandra Quave, senior author of the study and associate professor in Emory School of Medicine’s Department of Dermatology and the Center for the Study of Human Health. “Our findings open a new potential approach to deal with these infections, including those caused by deadly Candida auris.”
C. auris is often multidrug-resistant and has a high mortality rate, leading the Centers for Disease Control and Prevention (CDC) to label it a serious global health threat.
“It’s a really bad bug,” says Lewis Marquez, first author of the study and a graduate student in Emory’s molecular systems and pharmacology program. “Between 30 to 60% of the people who get infected with C. auris end up dying.”
An emerging threat
Candida is a yeast often found on the skin and in the digestive tract of healthy people. Some species, such as Candida albicans, occasionally grow out of control and cause mild infections in people. In more serious cases, Candida can invade deep into the body and cause infections in the bloodstream or organs such as the kidney, heart or brain.
Immunocompromised people, including many hospital patients, are most at risk for invasive Candida infections, which are rapidly evolving drug resistance.
In 2007, the new Candida species, C. auris, emerged in a hospital patient in Japan. Since then, C. auris has caused health care-associated outbreaks in more than a dozen countries around the world with more than 3,000 clinical cases reported in the United States alone.
A ‘natural’ approach to drug discovery
Quave is an ethnobotanist, studying how traditional people have used plants for medicine to search for promising new candidates for modern-day drugs. Her lab curates the Quave Natural Product Library, which contains 2,500 botanical and fungal natural products extracted from 750 species collected at sites around the world.
“We’re not taking a random approach to identify potential new antimicrobials,” Quave says. “Focusing on plants used in traditional medicines allows us to hone in quickly on bioactive molecules.”
Previously, the Quave lab had found that the berries of the Brazilian peppertree, a plant used by traditional healers in the Amazon for centuries to treat skin infections and some other ailments, contains a flavone-rich compound that disarms drug-resistant staph bacteria. Screens by the Quave lab had also found that the leaves of the Brazilian peppertree contain PGG, a compound that has shown antibacterial, anticancer and antiviral activities in previous research.
A 2020 study by the Quave lab, for instance, found that PGG inhibited growth of Carbapenem-resistant Acinetobacter baumannii, a bacterium that infects humans and is categorized as one of five urgent threats by the CDC.
The Brazilian peppertree, an invasive weed in Florida, is a member of the poison ivy family. “PGG has popped up repeatedly in our laboratory screens of plant compounds from members of this plant family,” Quave says. “It makes sense that these plants, which thrive in really wet environments, would contain molecules to fight a range of pathogens.”
Experimental results
The Quave lab decided to test whether PGG would show antifungal activity against Candida. Laboratory-dish experiments demonstrated that PGG blocked around 90% of the growth in 12 strains from four species of Candida: C. albicans, multidrug-resistant C. auris and two other multidrug-resistant non-albicans Candida species.
PGG is a large molecule known for its iron-binding properties. The researchers tested the role of this characteristic in the antifungal activity.
“Each PGG molecule can bind up to five iron molecules,” Marquez explains. “When we added more iron to a dish, beyond the sequestering capacity of the PGG molecules, the fungi once again grew normally.”
Dish experiments also showed that PGG was well-tolerated by human kidney, liver and epithelial cells. “Iron in human cells is generally not free iron,” Marquez says. “It is usually bound to a protein or is sequestered inside enzymes.”
A potential topical treatment
Previous animal studies on PGG have found that the molecule is metabolized quickly and removed from the body. Instead of an internal therapy, the researchers are investigating its potential efficacy as a topical antifungal.
“If a Candida infection breaks out on the skin of a patient where a catheter or other medical instrument is implanted, a topical antifungal might prevent the infection from spreading and entering into the body,” Marquez says.
As a next step, the researchers will test PGG as a topical treatment for fungal skin infections in mice.
Meanwhile, Quave and Marquez have applied for a provisional patent for the use of PGG for the mitigation of fungal infections.
“These are still early days in the research, but another idea that we’re interested in pursuing is the potential use of PGG as a broad-spectrum microbial,” Quave says. “Many infections from acute injuries, such as battlefield wounds, tend to be polymicrobial so PGG could perhaps make a useful topical treatment in these cases.”
Scientists from the University of Toronto are co-authors of the paper, including Yunjin Lee, Dustin Duncan, Luke Whitesell and Leah Cowen. Whitesell and Cowen are co-founders and shareholders in Bright Angel Therapeutics, a platform company for development of antifungal therapeutics, and Cowen is a science advisor for Kapoose Creek, a company that harnesses the therapeutic potential of fungi.
The work was supported by grants from the National Institutes of Health, National Center for Complementary and Integrative Health; the Jones Center at Ichauway, the CIHR Frederick Banting and Charles Best Canada Graduate Scholarship and the Canadian Institutes of Health Research Foundation.
Related
Extracts from two wild plants inhibit COVID virus, study finds
Scientists identify chemicals in noxious weed that 'disarm' deadly bacteria
Friday, September 8, 2023
NIH funds Emory center to advance cellular mechanics
By Carol Clark
The National Institutes of Health (NIH) awarded Emory University $5.6 million to establish a national center to advance pioneering technology for cellular mechanics. The center is directed by Khalid Salaita, Emory professor of chemistry, whose lab developed the first sensors for detecting cell-receptor forces at the molecular level.
“We’ve been working on our molecular-force probes for more than a decade,” Salaita says. “We’ve demonstrated that these probes can be used to visualize, measure and map cellular forces down to the level of piconewtons. The center allows us to get this technology into the hands of end users — researchers in the biomedical sciences.”
The Center for Molecular Mechanobiology encompasses labs from seven leading research institutions including: Children’s Hospital of Philadelphia, Dana-Farber Cancer Institute, Emory, Georgia Tech, Memorial Sloan Kettering, University of Utah and Vanderbilt University.
The center members will use the molecular-force probes to investigate the biomechanics of processes such as the clotting of blood cells, the response of immune cells to an infection and the migration of cancer cells. Better understanding these processes may lead to the development of new treatments and therapies for a range of diseases and disorders.
In addition to supplying the technology, the center will train researchers to use the molecular-force probes and help adapt the technology to answer specific biomedical research questions.
“Working directly with the research community will help us to further refine and optimize the technology,” Salaita says. “We envision that measuring cellular forces will soon become part of the standard repertoire of biochemical techniques that scientists use to study living systems.”
The center’s associate directors are Yonggan Ke (associate professor in the Wallace H. Coulter Department of Biomedical Engineering at Emory and Georgia Tech) and Alexa Mattheyses (associate professor in the Department of Cell Developmental and Integrative Biology at the University of Alabama).
The five-year award from the National Institute of General Medical Sciences is part of the NIH Biomedical Technology Optimization and Dissemination Centers program. The goal is to optimize and disseminate state-of-the-art, late-stage biomedical technologies.
The first detailed view of mechanical forces
The Salaita lab works at the intersection of chemistry, biology and the physical sciences. It uses the building blocks of nature — nucleic acids — to create synthetic micro motors and probes for investigating fundamental questions of biology.
The molecular-force probes, developed by the Salaita lab in 2011, provide the first detailed view of the mechanical forces on the surface of a cell. The technology can detect mechanical forces as fleeting as the blink of an eye and as faint as piconewtons — about one billionth the weight of a paperclip.
The probes are made from strands of synthetic DNA tagged with fluorescence so that they function like molecular beacons, shining when they sense force. The technique is noninvasive, does not modify the cell and can be done with a standard fluorescence microscope.
In 2014, the lab used the new method to demonstrate how adherent cells — the kind that form the architecture of all multicellular organisms — mechanically sense their environments, migrate and stick to things.
In 2016, the molecular-force probes provided the first direct evidence for the mechanical forces of T cells, the security guards of the immune system. The lab’s experiments on T cells drawn from mice showed how they use a kind of mechanical “handshake” to test whether a cell they encounter is a friend or a foe.
In 2017, the lab shined its molecular beacons on platelets, the cells in the blood whose job is to stop bleeding by sticking together to form clots and plug up a wound. That work revealed the key molecular forces on platelets that trigger the clotting process.
In 2020, the lab and its collaborators combined advances in optical imaging with the molecular-force probes to capture forces at a resolution of 25 nanometers — far shorter than the length of a light wave. “That resolution is akin to being on the moon and seeing the ripples caused by raindrops hitting the surface of a lake on the Earth,” Salaita said at the time.
Key technological goals
The Center for Molecular Mechanobiology will build on this foundational work of the Salaita lab. It will focus on three key technological development goals:
• Optimizing the highest-resolution technique of the molecular-force probes so that it can be applied to a range of research questions.
• Tagging cells based on their force level in order to use force as a marker to barcode cells and their receptors. The idea is to classify the mechanics of individual cells and then link these classifications to gene-expression levels to study the cause-and-effect relationships.
• Amplifying the molecular-force signals to better understand the role of even the weakest forces involved in cellular mechanics, including those involved in the immune response.
Researchers from throughout the country will come to the Center for Molecular Mechanobiology to receive hands-on training in the molecular-force probes and then return to their home labs to become ambassadors for the technology.
“We’ll be adding a whole other layer of information for researchers working on everything from designing vaccines to cancer immunotherapy agents,” Salaita says.
Decades ago, he points out, complicated techniques such as crystallography, PCR and mass spectrometry were not frequently used but have since become routine workhorses in the biomedical sciences.
“We are catalyzing the process of spreading our technology so that studying biomechanics also becomes common and routine in biology,” Salaita says. “Molecular forces are a missing piece to understanding the way biology works.”
Related:
‘Firefly’ imaging method makes cellular forces visible
Chemists reveal the force within you
T cells use ‘handshakes’ to sort friends from foes
New methods reveal the mechanics of blood clotting
@font-face
{font-family:"Cambria Math";
panose-1:2 4 5 3 5 4 6 3 2 4;
mso-font-charset:0;
mso-generic-font-family:roman;
mso-font-pitch:variable;
mso-font-signature:-536870145 1107305727 0 0 415 0;}@font-face
{font-family:Calibri;
panose-1:2 15 5 2 2 2 4 3 2 4;
mso-font-charset:0;
mso-generic-font-family:swiss;
mso-font-pitch:variable;
mso-font-signature:-536859905 -1073732485 9 0 511 0;}p.MsoNormal, li.MsoNormal, div.MsoNormal
{mso-style-unhide:no;
mso-style-qformat:yes;
mso-style-parent:"";
margin:0in;
mso-pagination:widow-orphan;
font-size:12.0pt;
font-family:"Calibri",sans-serif;
mso-ascii-font-family:Calibri;
mso-ascii-theme-font:minor-latin;
mso-fareast-font-family:Calibri;
mso-fareast-theme-font:minor-latin;
mso-hansi-font-family:Calibri;
mso-hansi-theme-font:minor-latin;
mso-bidi-font-family:"Times New Roman";
mso-bidi-theme-font:minor-bidi;}a:link, span.MsoHyperlink
{mso-style-priority:99;
color:#0563C1;
mso-themecolor:hyperlink;
text-decoration:underline;
text-underline:single;}a:visited, span.MsoHyperlinkFollowed
{mso-style-noshow:yes;
mso-style-priority:99;
color:#954F72;
mso-themecolor:followedhyperlink;
text-decoration:underline;
text-underline:single;}.MsoChpDefault
{mso-style-type:export-only;
mso-default-props:yes;
font-family:"Calibri",sans-serif;
mso-ascii-font-family:Calibri;
mso-ascii-theme-font:minor-latin;
mso-fareast-font-family:Calibri;
mso-fareast-theme-font:minor-latin;
mso-hansi-font-family:Calibri;
mso-hansi-theme-font:minor-latin;
mso-bidi-font-family:"Times New Roman";
mso-bidi-theme-font:minor-bidi;
mso-font-kerning:0pt;
mso-ligatures:none;}div.WordSection1
{page:WordSection1;}
Thursday, August 10, 2023
Images of enzyme in action reveal secrets of antibiotic-resistant bacteria
By Carol Clark
Bacteria draw from an arsenal of weapons to combat the drugs intended to kill them. Among the most prevalent of these weapons are ribosome-modifying enzymes. These enzymes are growing increasingly common, appearing worldwide in clinical samples in a range of drug-resistant bacteria.
Now scientists have captured the first images of one important class of these enzymes in action. The images show how the enzymes latch onto a particular site on the bacterial ribosome and squeeze it like a pair of tweezers to extract an RNA nucleotide and alter it.
The Proceedings of the National Academy of Sciences (PNAS) published the findings, led by scientists at Emory University. The advanced technique of cryoelectron microscopy made the ultra-high-resolution, three-dimensional snapshots possible.
“Seeing is believing,” says Christine Dunham, Emory professor of chemistry and co-corresponding author of the paper. “The minute you see biological structures interacting in real life at the atomic level it’s like solving a jigsaw puzzle. You see how everything fits together and you get a clearer idea of how things work.”
The insights may lead to the design of new antibiotic therapies to inhibit the drug-resistance activities of RNA methyltransferase enzymes. These enzymes transfer a small hydrocarbon known as a methyl group from one molecule to another, a process known as methylation.
“Methylation is one of the smallest chemical modifications in biology,” says Graeme Conn, professor of biochemistry in Emory’s School of Medicine and co-corresponding author of the paper. “But this tiny modification can fundamentally change biology. In this case, it confers resistance that allows bacteria to evade an entire class of antibiotics.”
Both Conn and Dunham are also members of the Emory Antibiotic Resistance Center.
First author of the paper is Pooja Srinivas, who did the work as a PhD candidate in Emory’s graduate program in molecular and systems pharmacology. She has since graduated and is now a postdoctoral fellow at the University of Washington.
Understanding the ribosome
Dunham is a leading expert on the ribosome — an elaborate structure that operates like a factory within a cell to manufacture proteins. Proteins are the machines that make cells run while nucleic acids such as DNA and RNA store the blueprints for life. The ribosome is made mostly of RNA, which does not just store information but can also act as an enzyme, catalyzing chemical reactions.
One goal of Dunham’s lab is to find ways to manipulate bacterial ribosomes to make them more susceptible to antimicrobials. If an antimicrobial successfully inactivates bacterial ribosomes, that shuts down the manufacturing of proteins essential for bacterial growth and survival.
The idea is to exploit differences in human cellular ribosomes and bacterial ribosomes, so that only the bacteria is targeted by an antimicrobial drug.
Antimicrobials, however, need to get past bacterial defenses.
“It’s like a molecular arms race,” Dunham explains. Bacteria keep evolving new weapons as a defense against drugs, while scientists evolve new strategies to disarm bacteria.
Enzymes that modify the ribosome
Conn is a leading expert in the bacterial defense weapons known as ribosomal RNA methyltransferase enzymes. This family of enzymes was originally discovered in soil bacteria. They are now increasingly found in bacterial infections in people and animals, making these infections harder to treat.
“They keep turning up more and more often in clinical samples of some nasty bacterial pathogens in different parts of the world,” Conn says.
The enzymes can drive deadly drug-resistance in pathogens such as E. coli, Salmonella, Klebsiella pneumoniae, Pseudomonas aeruginosa and Enterobacteriaceae. The enzymes add a methyl group at a specific site on the bacterial ribosome. That addition blocks the ability of a class of antibiotics known as aminoglycosides to bind and cause their antibacterial action.
For the PNAS paper, the researchers focused on a culprit within this family of enzymes known as ribosomal RNA methyltransferase C, or RmtC.
A complicated enzyme
For decades, researchers have relied on a technique known as X-ray crystallography to reveal the atomic details of how molecular machines work when the molecules are arranged in a crystal.
In 2015, for example, Dunham’s lab obtained precise pictures through X-ray crystallography of how an enzyme known as HigB rips up RNA to inhibit growth of the bacteria. By restraining the growth of the bacteria that makes it, HigB establishes a dormant “persister cell” state that makes the bacteria tolerant to antibiotics.
The secrets of how the RmtC enzyme interacts with the ribosome, however, eluded X-ray crystallography.
“RmtC is much more complicated,” Dunham explains. “It’s an interesting enzyme from a basic science perspective because it looks so different from others.”
A resolution revolution
Recent advances in cryoelectron microscopy opened the door to zooming in on the complex mechanisms of RmtC.
Cryoelectron microscopy does not require crystallization to reveal the structures of molecules and how they interact. Instead, liquid samples are frozen rapidly to form a glassy matrix. The glassy matrix retains the three-dimensional structure of molecules and protects them from deterioration by the intense electron beam.
Meisam Nosrati, a former postdoctoral fellow in the Conn lab and a co-author of the PNAS paper, prepared samples of RmtC interacting with part of an E. coli ribosome. He tapped the expertise of co-author Lindsay Comstock, a chemist at Wake Forest University who developed a technique to trap and stabilize the enzyme in the needed position.
Nosrati then froze the samples on a tiny grid and sent them to the Pacific Northwest Center for Cryo-EM for imaging.
As a graduate student in the Dunham lab, Pooja Srinivas then analyzed and interpreted the microscopy dataset. She used computer algorithms to stitch together thousands of individual images.
The result turned the images into a flipbook that revealed the complicated structure of RmtC in action.
“The enzyme latches on like a pincer to the ribosome,” Dunham explains. “It tightens its grip until it squeezes out a nucleotide from the interior of an RNA helix. It then chemically modifies that nucleotide.”
The enzyme is exquisitely specific about where it binds to the ribosome, a huge macromolecule made up of 50 different proteins and 6,000 different RNA nucleotides.
The researchers used biochemistry techniques to validate that what they observed matched previous findings for how RmtC makes bacteria resistant to aminoglycoside antimicrobials that target the ribosome.
Strategies for new therapies
The researchers are now trying to develop new ways to counter the effects of RmtC and related enzymes based on the new information.
“Knowledge of the shape of the enzyme as its performs its chemical reaction gives us new targets to inhibit its effects,” Conn says. “For instance, we could target the pincer action of the enzyme to try to prevent it from squeezing and binding to the ribosome. We now know that the enzyme forms a pocket on its surface where a small molecule might sit to block this action.”
Additional co-authors of the PNAS paper are Natalia Zelinskaya and Debayan Dey, research scientists in the Conn lab. Funding for the work was provided by the National Institutes of Health and the Burroughs Wellcome Fund Investigator in the Pathogenesis of Infectious Disease Award.
Related:
Biochemist Dunham shifts the frame on proteins
New molecule found in chestnut trees disarms dangerous staph bacteria
Wednesday, July 26, 2023
Merck Prize boosts work on air sensor for pandemic pathogens
Merck KGaA, Darmstadt, Germany, awarded its 2023 Future Insight Prize to Khalid Salaita, professor of chemistry at Emory University. The award comes with $540,000 to fund the next phase of research into an air sensor that can continuously monitor indoor spaces for pathogens that can cause pandemics.
“I’m extremely thankful to receive the Future Insight Prize as this enables us to continue our path toward an early-warning system for emerging threats,” Salaita says. “Our research sets the stage for fully automated detection of airborne pathogens without human intervention or sample processing.”
The Merck Future Insight Prize recognizes groundbreaking ideas to solve some of the world’s most pressing challenges in health, nutrition and energy.
The Salaita lab’s sensor, a rolling micro-motor called “Rolosense,” holds the potential to help mitigate, or even prevent, a pandemic.
Thursday, February 9, 2023
Extracts from two wild plants inhibit COVID-19 virus, study finds
By Carol Clark
Two common wild plants contain extracts that inhibit the ability of the virus that causes COVID-19 to infect living cells, an Emory University study finds. Scientific Reports published the results — the first major screening of botanical extracts to search for potency against the SARS-CoV-2 virus.
In laboratory dish tests, extracts from the flowers of tall goldenrod (Solidago altissima) and the rhizomes of the eagle fern (Pteridium aquilinum) each blocked SARS-CoV-2 from entering human cells.
The active compounds are only present in miniscule quantities in the plants. It would be ineffective, and potentially dangerous, for people to attempt to treat themselves with them, the researchers stress. In fact, the eagle fern is known to be toxic, they warn.
“It’s very early in the process, but we’re working to identify, isolate and scale up the molecules from the extracts that showed activity against the virus,” says Cassandra Quave, senior author of the study and associate professor in Emory School of Medicine’s Department of Dermatology and the Center for the Study of Human Health. “Once we have isolated the active ingredients, we plan to further test for their safety and for their long-range potential as medicines against COVID-19.”
A powerful tool for drug discovery
Quave is an ethnobotanist, studying how traditional people have used plants for medicine to identify promising new candidates for modern-day drugs. Her lab curates the Quave Natural Product Library, which contains thousands of botanical and fungal natural products extracted from plants collected at sites around the world.
 |
| Tall goldenrod |
Caitlin Risener, a PhD candidate in Emory’s Molecular and Systems Pharmacology graduate program and the Center for the Study of Human Health, is first author of the current paper.
In previous research to identify potential molecules for the treatment of drug-resistant bacterial infections, the Quave lab focused on plants that traditional people had used to treat skin inflammation.
Given that COVID-19 is a newly emerged disease, the researchers took a broader approach. They devised a method to rapidly test more than 1,800 extracts and 18 compounds from the Quave Natural Product Library for activity against SARS-CoV-2.
“We’ve shown that our natural products library is a powerful tool to help search for potential therapeutics for an emerging disease,” Risener says. “Other researchers can adapt our screening method to search for other novel compounds within plants and fungi that may lead to new drugs to treat a range of pathogens.”
Picking the locks on a cell’s surface
SARS-CoV-2 is an RNA virus with a spike protein that can bind to a protein called ACE2 on host cells. “The viral spike protein uses the ACE2 protein almost like a key going into a lock, enabling the virus to break into a cell and infect it,” Quave explains.
The researchers devised experiments with virus-like particles, or VLPs, of SARS-CoV-2, and cells programmed to overexpress ACE2 on their surface. The VLPs were stripped of the genetic information needed to cause a COVID-19 infection. Instead, if a VLP managed to bind to an ACE2 protein and enter a cell, it was programmed to hijack the cell’s machinery to activate a fluorescent green protein.
A plant extract was added to the cells in a petri dish before introducing the viral particles. By shining a fluorescent light on the dish, they could quickly determine whether the viral particles had managed to enter the cells and activate the green protein.
The researchers identified a handful of hits for extracts that protected against viral entry and then homed in on the ones showing the strongest activity: Tall goldenrod and eagle fern. Both plant species are native to North America and are known for traditional medicinal uses by Native Americans.
Additional experiments showed that the protective power of the plant extracts worked across four variants of SARS-CoV-2: Alpha, theta, delta and gamma.
Confirming the results with infectious virus
To further test these results, the Quave lab collaborated with co-author Raymond Schinazi, Emory professor of pediatrics, director of Emory’s Division of Laboratory of Biochemical Pharmacology and co-director of the HIV Cure Scientific Working Group within the NIH-sponsored Emory University Center for AIDS Research. A world leader in antiviral development, Schinazi is best known for his pioneering work on breakthrough HIV drugs.
The higher biosecurity rating of the Schinazi lab enabled the researchers to test the two plant extracts in experiments using infectious SARS-CoV-2 virus instead of VLPs. The results confirmed the ability of the tall goldenrod and eagle fern extracts to inhibit the ability of SARS-CoV-2 to bind to a living cell and infect it.
“Our results set the stage for the future use of natural product libraries to find new tools or therapies against infectious diseases," Quave says.
As a next step, the researchers are working to determine the exact mechanism that enables the two plant extracts to block binding to ACE2 proteins.
A hands-on connection to nature
For Risener, one of the best parts about the project is that she collected samples of tall goldenrod and eagle fern herself. In addition to gathering medicinal plants from around the globe, the Quave lab also makes field trips to the forests of the Joseph W. Jones Research Center in South Georgia. The Woodruff Foundation established the center to help conserve one of the last remnants of the unique longleaf pine ecosystem that once dominated the southeastern United States.
“It’s awesome to go into nature to identify and dig up plants,” Risener says. “That’s something that few graduate students in pharmacology get to do. I’ll be covered in dirt from head to toe, kneeling on the ground and beaming with excitement and happiness.”
She also assists in preparing the plant extracts and mounting the specimens for the Emory Herbarium. “When you collect a specimen yourself, and dry and preserve the samples, you get a personal connection,” she says. “It’s different from someone just handing you a vial of plant material in a lab and saying, ‘Analyze this.’”
After graduating, Risener hopes for a career in outreach and education for science policy surrounding research into natural compounds. A few of the more famous medicines derived from botanicals include aspirin (from the willow tree), penicillin (from fungi) and the cancer therapy Taxol (from the yew tree).
“Plants have such chemical complexity that humans probably couldn’t dream up all the botanical compounds that are waiting to be discovered,” Risener says. “The vast medicinal potential of plants highlights the importance of preserving ecosystems.”
Co-authors of the current paper include: Sumin Woo, Tharanga Samarakoon, Marco Caputo and Emily Edwards (the Quave lab and Emory’s Center for the Study of Human Health); Keivan Zandi, Shu Ling Goh and Jessica Downs-Bowen (the Schinazi lab); Kier Klepzig (Joseph W. Jones Research Center); and Wendy Applequist (Missouri Botanical Garden).
Funding for the paper was provided by the Marcus Foundation, the NIH-funded Center for AIDS Research and the NIH National Center for Complementary and Integrative Health.
Related:
Into the heart of brightness: An ethnobotanist's memoir
The plant hunters: Students search for secrets of ancient remedies
New molecule found in chestnut trees disarms dangerous bacteria